Marine bacteria have learned to degrade industrial pollutants, offering new hope for scientists struggling to clean up tainted ocean water. Human ingenuity has led to the development of chemical configurations that are new to nature. Unfortunately, these human-made compounds — which function as plasticisers, deadly nerve agents, pesticides, and flame retardants — accumulate in marine environments. New evidence shows that marine bacteria have evolved to degrade these compounds as an alternative source of phosphorus, an essential element found in DNA and RNA. By inserting genes from marine bacteria into E. coli, scientists were able to create “tiny pollution scavengers” in the lab.
Humankind is reshaping the oceans and seas: As temperatures rise, dissolved oxygen levels drop, and ecosystems are disrupted. But global warming is not our only assault on marine life: clever chemists have devised new classes of molecules that life on Earth has never encountered before. These industrial and agricultural pollutants further undermine the delicate communities of our blue planet. But with change comes opportunity, and an intriguing evolutionary question arises from this unfortunate situation: Has marine life adapted to human pollution and somehow turned the tables in their favour?
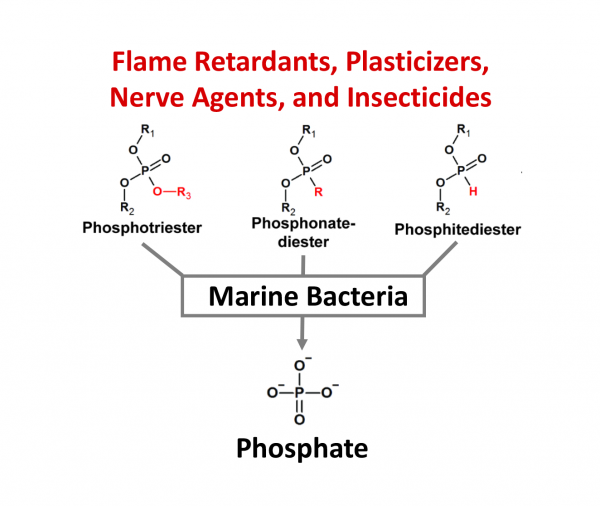
This is the question that inspired a team of scientists including Liam Longo, Specially Appointed Associate Professor of Earth-life Science Institute (ELSI) at Tokyo Institute of Technology, the late Prof. Dan Tawfik (Weizmann Institute of Science), Dr. Dragana Despotović (Weizmann Institute of Science), and Prof. Einat Segev (Weizmann Institute of Science) to embark on an interdisciplinary journey in search of new enzymes in the sea. The research centred on an incredible opportunity created by polluted oceans: In pristine waters, marine bacteria often struggle to find phosphorus, an essential element that is present in DNA. Because phosphorus is so important to life, there is stiff competition to absorb it. As it turns out, man-made pollutants are rich in phosphorus (Figure 1), though special enzymes are required to liberate it. While most researchers assumed that these enzymes are extremely rare, the Prof. Longo and his collaborators were sceptical.
To test their hypothesis, they collected bacterial samples from the Mediterranean Sea and the Red Sea. After isolating bacteria from the environment, they challenged it to grow on pollutants as the only source of phosphorus. And the bacteria grew! In fact, of the 15 different pollutants tested, the growth of at least one type of marine bacteria was observed in each case. The ability to degrade pollutants, therefore, is far more widespread than anyone imagined.
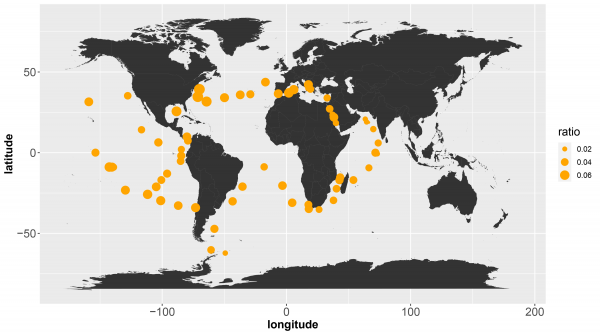
Next, the researchers identified two of the enzymes that made this new chemistry possible. Using the amino acid sequences of these new enzymes as a starting point, the genomes of all known marine bacteria could be analysed. Once again, the researchers observed that these enzymes are widespread, with sequence matches found in bacteria from all around the world (Figure 2).
Finally, by inserting a gene from the marine bacteria into E. coli, the scientists created a strain of E. coli that can degrade pollutants. This result demonstrates that these enzymes can be used to clean up contaminated waters by a process called bioremediation.
Does all of this good news mean that the problem of industrial pollution is solved? No. Industrial pollutants remain extremely toxic to sea life, especially sea animals, where they often act as neurotoxins. We must continue to fight for the decontamination of our oceans and seas.
| Journal | Proceedings of the National Academy of Sciences |
| Tile of the paper | Utilisation of diverse organophosphorus pollutants by marine bacteria |
| Authors | Dragana Despotović1*, Einav Aharon1, Olena Trofimyuk1, Artem Dubovetskyi1,2, Kesava Phaneendra Cherukuri1, Yacov Ashani1, Or Eliason3, Martin Sperfeld3, Haim Leader1, Andrea Castelli4, Laura Fumagalli4, Alon Savidor5, Yishai Levin5, Liam M. Longo6,7*, Einat Segev3*, and Dan S. Tawfik1 |
| Affiliations |
|
| DOI | 10.1073/pnas.2203604119 |
| Online published date | 2 August 2022 |
